AGRP
Asteraceae Genomic Research Platform
Welcome to AGRP!156
Asteraceae is the largest angiosperm family, with more than 1,700 genera and 25,000 species. Asteraceae have a strong adaptability to the environment and are widely distributed in almost all parts of the world. It contains a large number of edible, medicinal and ornamental plants, which are of great value to human life.
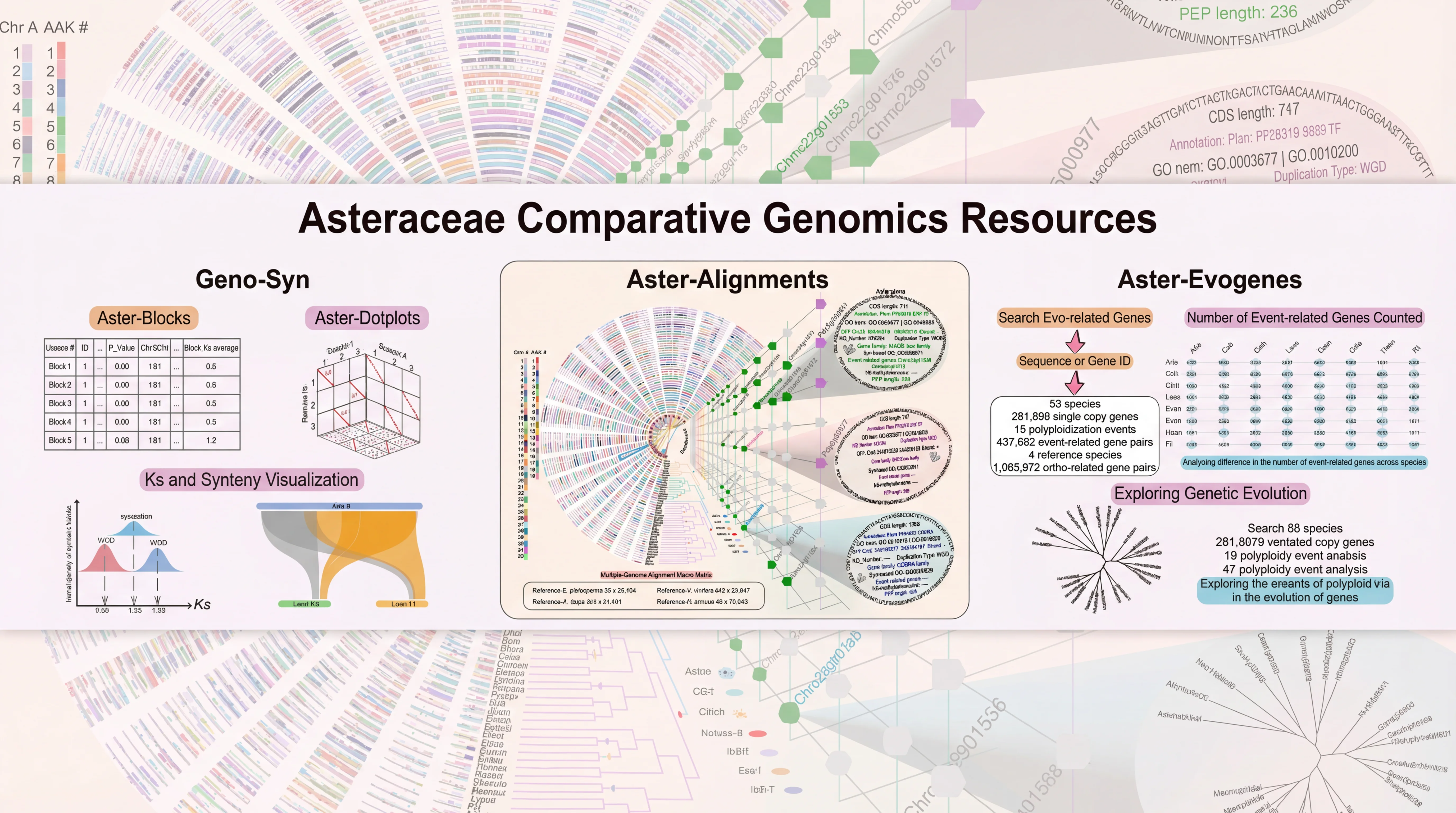
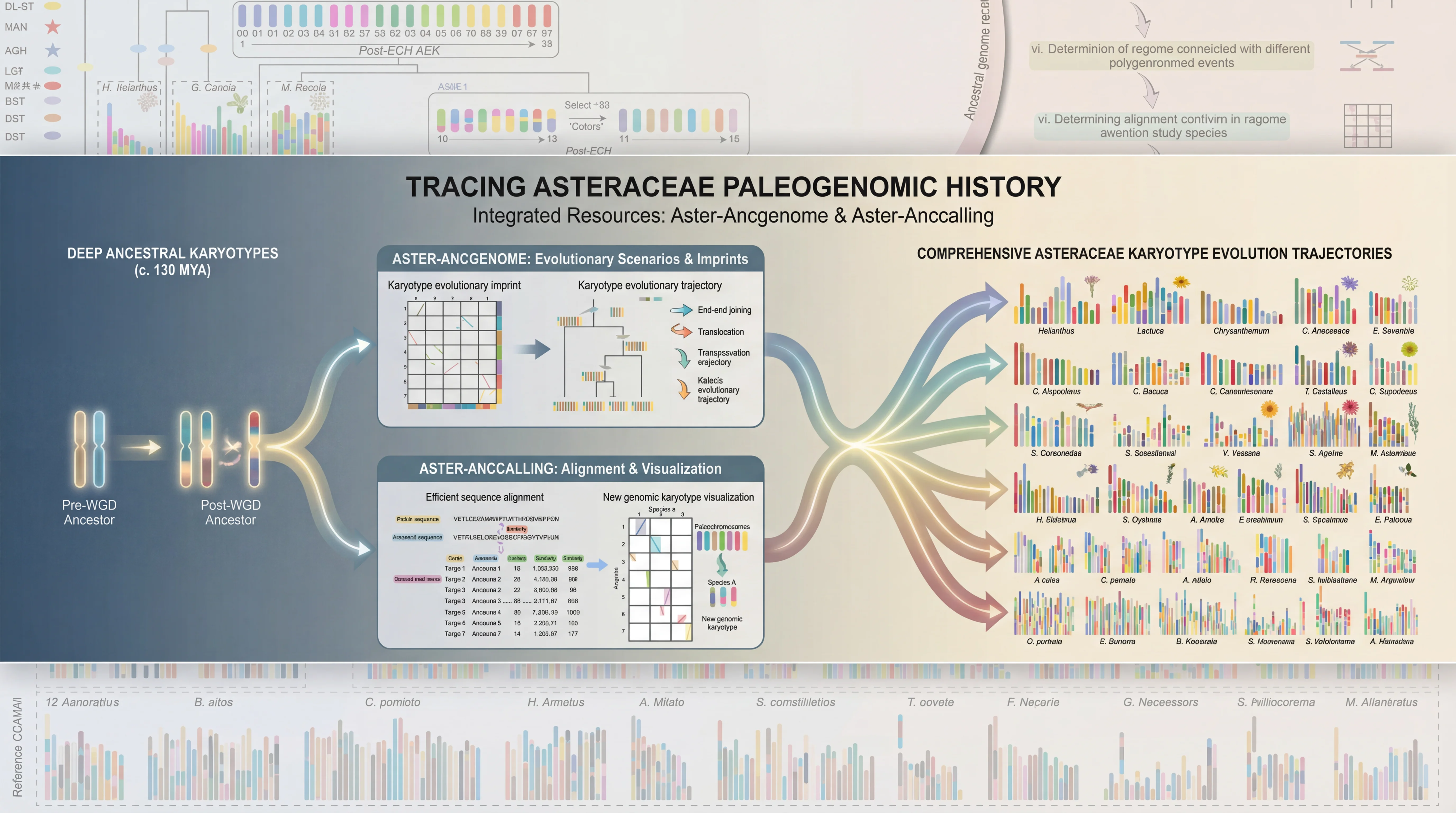
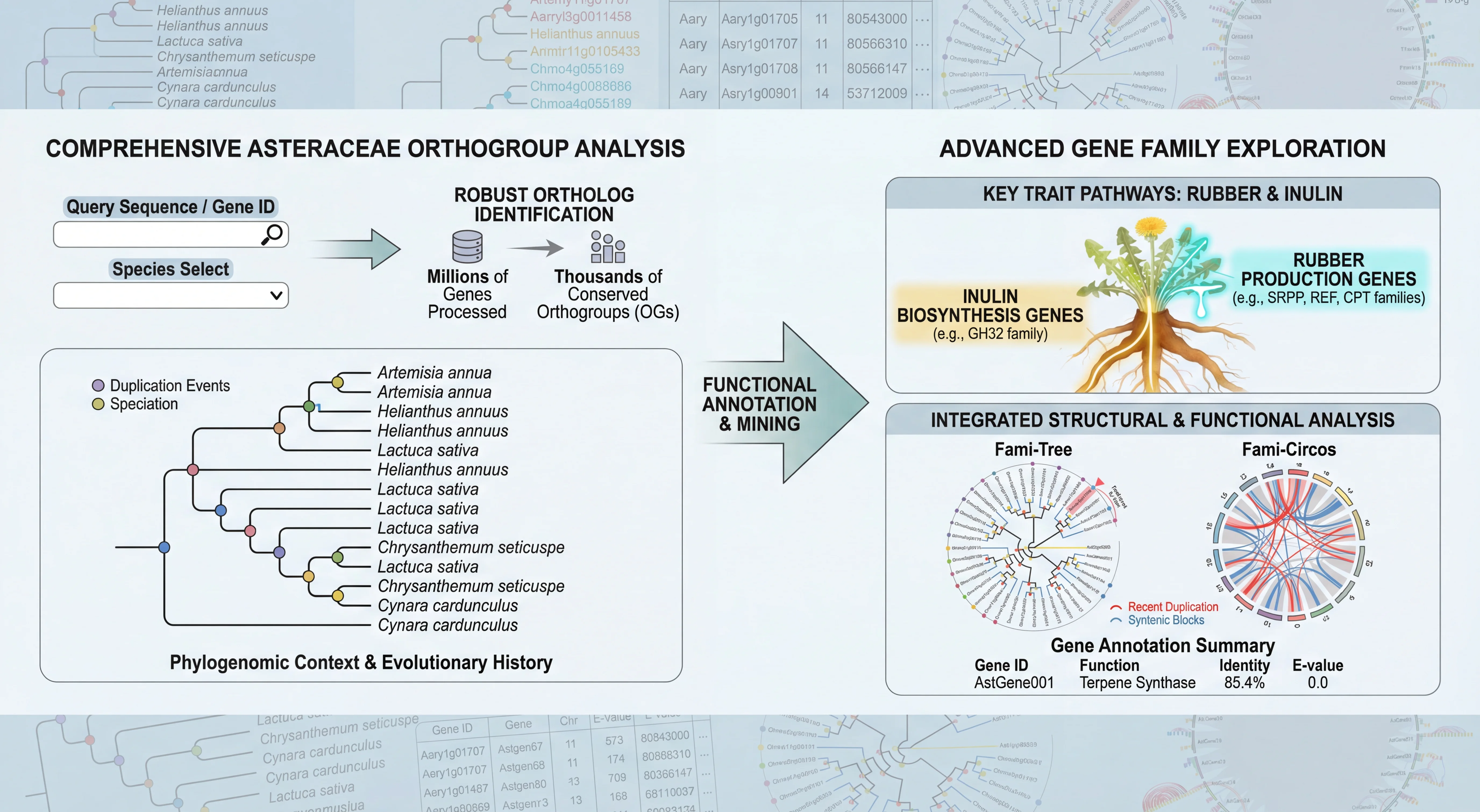
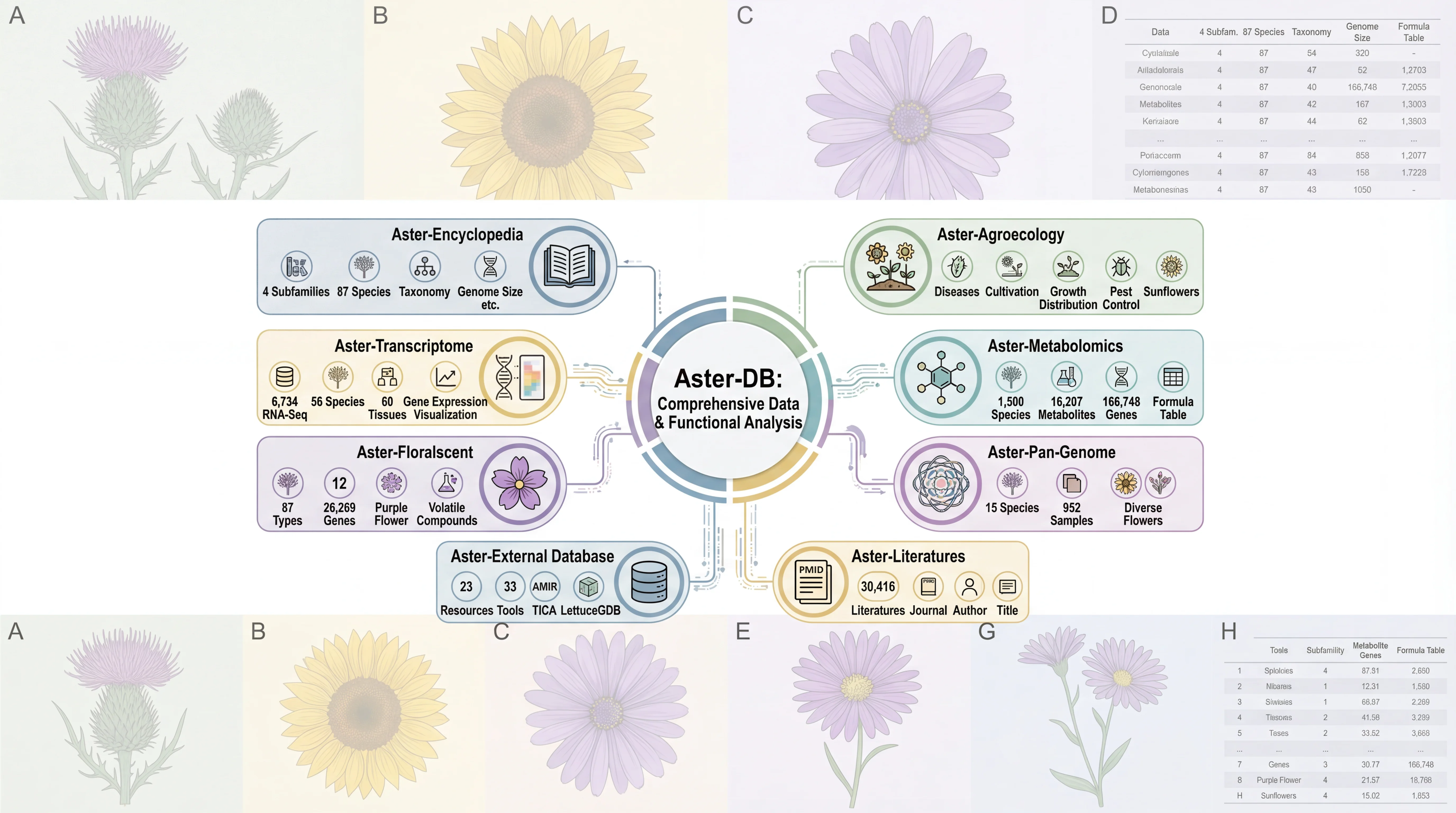
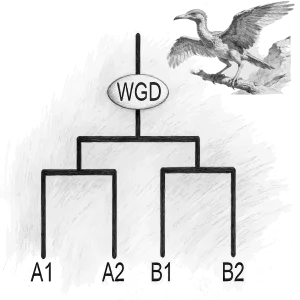
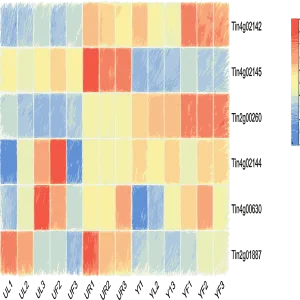
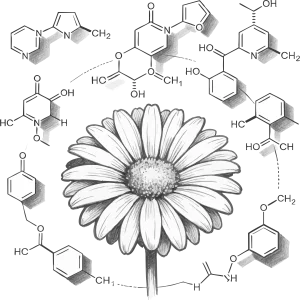
AGRP integrates 156 genomes, including 149 published genomes from the Asteraceae family, the newly sequenced canadensis genome, and 6 outgroup species. This platform provides annotation details for 3,137,213 protein-coding genes in Asteraceae genomes, along with resources for gene duplication types and synteny analysis; 180 trait-related gene families, including specialized inulin-related genes and rubber synthesis-related genes; the largest ancestral genome resource in Asteraceae, encompassing genomic data from 29 ancestral nodes; the most comprehensive collection of 6,734 transcriptomes in Asteraceae, along with 17,541 metabolite resources linked to 257,590 candidate genes; the first specialized floral scent metabolomics resource in Asteraceae, and encyclopedic resources of Asteraceae species. Interfaces for gene families and related analyses incorporate tools like Ortho-Tree, Fami-Tree, and Fami-Circos to support evolutionary gene studies. Additionally, the platform integrates 62 web-based analytical tools (34 developed in-house) and 74 downloadable tools, facilitating annotation, evolutionary analysis, and functional research for newly sequenced Asteraceae genomes. Beyond providing intuitive query, analysis, and visualization tools, it features a data submission portal, statistical charts, and user manuals to ensure accessibility for all users. AGRP simultaneously establishes a distinctive AGRP-Switch interface to explore various omics data resources within the AGRP database through an integrated approach, enabling users to perform one-stop searches.